Engineering base-excised aptamers for highly specific recognition of adenosine†
Abstract
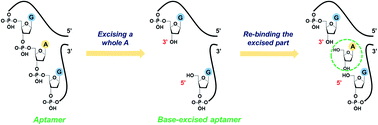
The DNA aptamer for adenosine and ATP has been used as a model system for developing analytical biosensors. For practical reasons, it is important to distinguish adenosine from ATP, although this has yet to be achieved despite extensive efforts made on selection of new aptamers. We herein report a strategy of excising an adenine nucleotide from the backbone of a one-site adenosine aptamer, and the adenine-excised aptamer allowed highly specific binding of adenosine. Cognate analytes including AMP, ATP, guanosine, cytidine, uridine, and theophylline all failed to bind to the engineered aptamer according to the SYBR Green I (SGI) fluorescence spectroscopy and isothermal titration calorimetry (ITC) results. Our A-excised aptamer has two binding sites: the original aptamer binding site in the loop and the newly created one due to base excision from the DNA backbone. ITC demonstrated that the A-excised aptamer strand can bind to two adenosine molecules, with a Kd of 14.8 ± 2.1 μM at 10 °C and entropy-driven binding. Since the wild-type aptamer cannot discriminate adenosine from AMP and ATP, we attributed this improved specificity to the excised site. Further study showed that these two sites worked cooperatively. Finally, the A-excised aptamer was tested in diluted fetal bovine serum and showed a limit of detection of 46.7 μM adenosine. This work provides a facile, cost-effective, and non-SELEX method to engineer existing aptamers for new features and better applications.


 Please wait while we load your content...
Please wait while we load your content...