Scaffold-based molecular design with a graph generative model†
Abstract
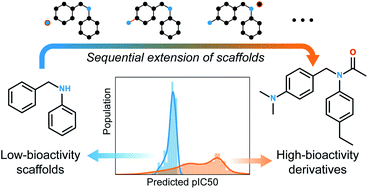
Searching for new molecules in areas like drug discovery often starts from the core structures of known molecules. Such a method has called for a strategy of designing derivative compounds retaining a particular scaffold as a substructure. On this account, our present work proposes a graph generative model that targets its use in scaffold-based molecular design. Our model accepts a molecular scaffold as input and extends it by sequentially adding atoms and bonds. The generated molecules are then guaranteed to contain the scaffold with certainty, and their properties can be controlled by conditioning the generation process on desired properties. The learned rule of extending molecules can well generalize to arbitrary kinds of scaffolds, including those unseen during learning. In the conditional generation of molecules, our model can simultaneously control multiple chemical properties despite the search space constrained by fixing the substructure. As a demonstration, we applied our model to designing inhibitors of the epidermal growth factor receptor and show that our model can employ a simple semi-supervised extension to broaden its applicability to situations where only a small amount of data is available.
- This article is part of the themed collection: Celebrating the 75th Anniversary of the Korean Chemical Society (KCS)


 Please wait while we load your content...
Please wait while we load your content...