Physicochemical interaction of cerium oxide nanoparticles with simulated biofluids, hemoglobin, insulin, and ds-DNA at 310.15 K†
Abstract
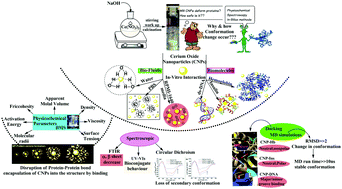
Cerium oxide nanoparticles (CNPs) are used as potential therapeutic nanomaterials to cure neurological oxidative diseases. This curing activity of CNPs has prompted the study of molecular interactions with several biologically significant molecules to probe conformational changes. CNPs@20 nm were synthesised and their structure confirmed by powder X-ray diffraction (XRD), transmission electron microscopy (TEM), X-ray photoelectron spectroscopy (XPS), Fourier-transform infrared spectroscopy (FTIR) and UV-vis spectrophotometry. In vitro interactions of 25–150 μM CNPs with water, phosphate buffered saline (PBS), and RPMI 1640 medium as simulated biofluids, and insulin (Ins) hemoglobin (Hb) and dsDNA at 37 °C were studied. Their physicochemical properties in terms of density (ρ, 103 kg m−3), viscosity (η, 10−3 kg m−1 s−1), surface tension (γ, mN m−1), friccohesity (σ, s m−1), activation energy (Δμ2* kJ mol−1) and molecular radii (r, nm) were measured using a density meter and Borosil Mansingh Survismeter (BMS), respectively. The data predicted a 1st order interaction and structure breaking behaviour of the CNPs with Hb, Ins, and dsDNA. Zeta potential measurements in biological media showed a net increase in negative charge magnitude with a higher colloidal stability. UV-vis and Fourier-transform infrared (FTIR) spectroscopic studies exhibited the biocompatibility of CNPs, with the sustaining of the functionality of native Hb, Ins, and dsDNA. A Kb value of 3.2 × 103 M−1 for CNPs–dsDNA implied its noncovalent interactions. A secondary conformational study using circular dichroism and FTIR revealed a decrease in the number of α helices and an increase in the number of β sheets. An in silico study was carried out via molecular docking and molecular dynamics simulations. Binding energies of −92.25, −91.36 and −106.84 kcal mol−1 were calculated for CNP–Hb, CNP–Ins, CNP–DNA docked complexes. The docking data show that CNP interactions via hydrogen bonding between Ins and Hb through neutral, polar and neutral, and nonpolar amino acids, respectively. The CNPs show electrostatic interactions at the minor groove of DNA. The study reports the unfolding mechanism investigated via physicochemical, spectroscopic and in silico approach of protein on interacting with CNPs, to guide their safer use in medicine and pharmaceutics.


 Please wait while we load your content...
Please wait while we load your content...