Single-molecule mechanical unfolding experiments reveal a critical length for the formation of α-helices in peptides†
Abstract
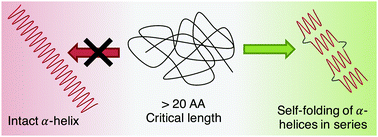
α-Helix is the most predominant secondary structure in proteins and supports many functions in biological machineries. The conformation of the helix is dictated by many factors such as its primary sequence, intramolecular interactions, or the effect of the close environment. Several computational studies have proposed that there is a critical maximum length for the formation of intact compact helical structures, supporting the fact that most intact α-helices in proteins are constituted of a small number of amino acids. To obtain a detailed picture on the formation of α-helices in peptides and their mechanical stability, we have synthesized a long homopolypeptide of about 90 amino acids, poly(γ-benzyl-L-glutamate), and investigated its mechanical behaviour by AFM-based single-molecule force spectroscopy. The characteristic plateaus observed in the force–extension curves reveal the unfolding of a series of small helices (from 1 to 4) of about 20 amino acid residues connected to each other, rather than a long helix of 90 residues. Our results suggest the formation of a tertiary structure made of short helices with kinks, instead of an intact compact helical structure for sequences of more than 20 amino acid residues. To our knowledge, this is the first experimental evidence supporting the concept of a helical critical length previously proposed by several computational studies.


 Please wait while we load your content...
Please wait while we load your content...