Repurposing drugs against the main protease of SARS-CoV-2: mechanism-based insights supported by available laboratory and clinical data†
Abstract
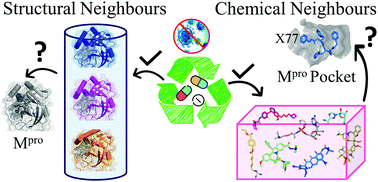
The ongoing global pandemic of COVID-19 has brought life to almost a standstill with the implementation of lockdowns and social distancing as some of the preventive measures in the absence of any approved specific therapeutic interventions. To combat this crisis, research communities worldwide are falling back on the existing repertoire of approved/investigational drugs to probe into their anti-coronavirus properties. In this report, we describe our unique efforts in identifying potential drugs that could be repurposed against the main protease of SARS-CoV-2 (SARS-CoV-2 Mpro). To achieve this goal, we have primarily exploited the principles of ‘neighbourhood behaviour’ in the protein 3D (workflow-I) and chemical 2D structural space (workflow-II) coupled with docking simulations and insights into the possible modes of action of the selected candidates from the available literature. This integrative approach culminated in prioritizing 29 potential repurpose-able agents (20 approved drugs and 9 investigational molecules) against SARS-CoV-2 Mpro. Apart from the approved/investigational anti-viral drugs, other notable hits include anti-bacterial, anti-inflammatory, anti-cancer and anti-coagulant drugs. Our analysis suggests that some of these drugs have the potential to simultaneously modulate the functions of viral proteins and the host response system. Interestingly, many of these identified candidates (12 molecules from workflow-I and several molecules, belonging to the chemical classes of alkaloids, tetracyclines, peptidomimetics, from workflow-II) are suggested to possess anti-viral properties, which is supported by laboratory and clinical data. Furthermore, this work opens a new avenue of research to probe into the molecular mechanism of action of many drugs, which are known to demonstrate anti-viral activity but are so far not known to target viral proteases.
- This article is part of the themed collection: Coronavirus articles - free to access collection


 Please wait while we load your content...
Please wait while we load your content...