Computational screening for nested organic cage complexes†
Abstract
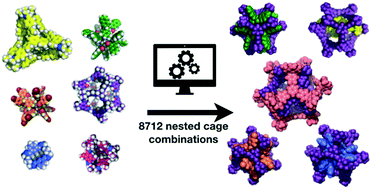
Supramolecular self-assembly has allowed the synthesis of beautiful and complex molecular architectures, such as cages, macrocycles, knots, catenanes, and rotaxanes. We focus here on porous organic cages, which are molecules that have an intrinsic cavity and multiple windows. These cages have been shown to be highly effective at molecular separations and encapsulations. We investigate the possibility of complexes where one cage sits within the cavity of another. We term this a ‘nested cage’ complex. The design of such complexes is highly challenging, so we use computational screening to explore 8712 different pair combinations, running almost 0.5 million calculations to sample the phase space of the cage conformations. Through analysing the binding energies of the assemblies, we identify highly energetically favourable pairs of cages in nested cage complexes. The vast majority of the most favourable complexes include the large imine cage reported by Gawroński and co-workers using a [8 + 12] reaction of 4-tert-butyl-2,6-diformylphenol and cis,cis-1,3,5-triaminocyclohexane. The most energetically favourable nested cage complex combines the Gawroński cage with a dodecaamide cage that has six vertices, which can sit in the six windows of the larger cage. We also identify cages that have favourable binding energies for self-catenation.
- This article is part of the themed collection: MSDE Emerging Investigators 2020


 Please wait while we load your content...
Please wait while we load your content...