Structural changes in the model of the outer cell membrane of Gram-negative bacteria interacting with melittin: an in situ spectroelectrochemical study
Abstract
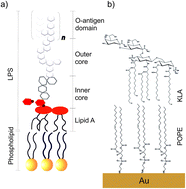
The cell membrane of Gram-negative bacteria interacting with an antimicrobial peptide presents a complex supramolecular assembly. Fabrication of models of bacterial cell membranes remains a large experimental challenge. Langmuir–Blodgett and Langmuir–Schaefer (LS–LB) transfer makes possible the deposition of multicomponent asymmetric lipid bilayers onto a gold surface. Two lipids: 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine (POPE) and di[3-deoxy-D-manno-octulosonyl]-lipid A (KLA) were used to deposit a model of the outer membrane of Gram-negative bacteria on the Au(111) substrate. The use of gold as the solid substrate enables control of the membrane potential. Molecular scale changes in the model membrane exposed to physiological electric fields and interacting with melittin antimicrobial peptide are discussed in this paper. The interaction of the outer membrane with melittin leads to an increase in the membrane capacitance and permeability to ions and water. The stability of the outer membrane with bound melittin decreases at positive membrane potentials. In situ polarization modulation infrared reflection absorption spectroscopy is used to investigate membrane potential-dependent changes in the structure of the outer membrane interacting with melittin. The hydration of the ester carbonyl groups is not affected by the interaction with melittin. However, the orientation and hydrogen bond network with the carboxylate groups in KLA changes drastically after POPE–KLA bilayer interacts with melittin. We propose that the positively charged groups in the amino acids present at the C-terminus of the peptide interact directly with the polar head group of KLA. Simultaneously, the packing order in hydrocarbon chains in the membrane with bound melittin increases. A hydrophobic match between the chains in the lipids and the peptide, which spans the membrane, seems to be responsible for the ordering of the hydrocarbon chains region of the bilayer. The N-terminus enters into the hydrophobic region of the membrane and forms a channel to the hydrophilic head groups in POPE.
- This article is part of the themed collection: Peptide-membrane interactions


 Please wait while we load your content...
Please wait while we load your content...