Single-molecule methods in structural DNA nanotechnology
Abstract
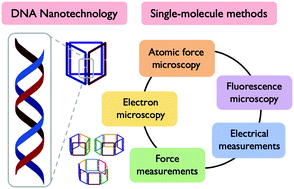
Single molecules can now be visualised with unprecedented precision. As the resolution of single-molecule experiments improves, so too does the breadth, quantity and quality of information that can be extracted using these methodologies. In the field of DNA nanotechnology, we use programmable interactions between nucleic acids to generate complex, multidimensional structures. We can use single-molecule techniques – ranging from electron and fluorescence microscopies to electrical and force spectroscopies – to report on the structure, morphology, robustness, sample heterogeneity and other properties of these DNA nanoconstructs. In this Tutorial Review, we will detail how complementarity between static and dynamic single-molecule techniques can provide a unified image of DNA nanoarchitectures. The single-molecule methods that we discuss provide unprecedented insight into chemical and structural behaviour, yielding not just an average outcome but reporting on the distribution of values, ultimately showing how bulk properties arise from the collective behaviour of individual structures. As the fields of both DNA nanotechnology and single-molecule characterisation intertwine, a feedback loop is generated between disciplines, providing new opportunities for the development and operation of DNA-based materials as sensors, delivery vehicles, machinery and structural scaffolds.


 Please wait while we load your content...
Please wait while we load your content...