The protein–water nuclear Overhauser effect (NOE) as an indirect microscope for molecular surface mapping of interaction patterns†
Abstract
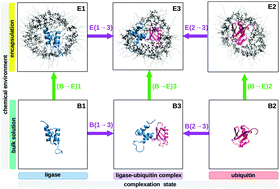
In this computational study, the intermolecular solute–solvent Nuclear Overhauser Effect (NOE) of the model protein ubiquitin in different chemical environments (free, bound to a partner protein and encapsulated) is investigated. Short-ranged NOE observables such as the NOE/ROE ratio reveal hydration phenomena on absolute timescales such as fast hydration sites and slow water clefts. We demonstrate the ability of solute–solvent NOE differences measured of the same protein in different chemical environments to reveal hydration changes on the relative timescale. The resulting NOE/ROE-surface maps are shown to be a central key for analyzing biologically relevant chemical influences such as complexation and confinement: the presence of a complexing macromolecule or a confining surface wall modulates the water mobility in the vicinity of the probe protein, hence revealing which residues of said protein are proximate to the foreign interface and which are chemically unaffected. This way, hydration phenomena can serve to indirectly map the precise influence (position) of other molecules or interfaces onto the protein surface. This proposed one-protein many-solvents approach may offer experimental benefits over classical one-protein other-protein pseudo-intermolecular transient NOEs. Furthermore, combined influences such as complexation and confinement may exert non-additive influences on the protein compared to a reference state. We offer a mathematical method to disentangle the influence of these two different chemical environments.


 Please wait while we load your content...
Please wait while we load your content...