Increasing the performance, trustworthiness and practical value of machine learning models: a case study predicting hydrogen bond network dimensionalities from molecular diagrams†‡
Abstract
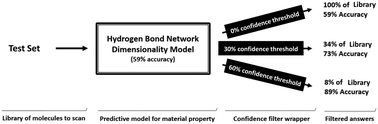
The performance of a model is dependent on the quality and information content of the data used to build it. By applying machine learning approaches to a standard chemical dataset, we developed a 4-class classification algorithm that is able to predict the hydrogen bond network dimensionality that a molecule would adopt in its crystal form with an accuracy of 59% (in comparison to a 25% random threshold), exclusively from two and lower dimensional molecular descriptors. Although better than random, the performance level achieved by the model did not meet the standards for its reliable application. The practical value of our model was improved by wrapping the model around a confidence tool that increases model robustness, quantifies prediction trust, and allows one to operate a classifier virtually up to any accuracy level. Using this tool, the performance of the model could be improved up to 73% or 89% with the compromise that only 34% and 8% of the total set of test examples could be predicted. We anticipate that the ability to adjust the performance of reliable 2D based models to the requirements of its different applications may increase their practical value, making them suitable to tasks that range from initial virtual library filtering to profile specific compound identification.
- This article is part of the themed collections: Introducing the CrystEngComm Advisory Board and their research and The Cambridge Structural Database - A wealth of knowledge gained from a million structures


 Please wait while we load your content...
Please wait while we load your content...