Label-free amplified fluorescence detection of DNA biomarkers based on KFP polymerase-driven double strand displacement reactions and magnetic nanoprobes†
Abstract
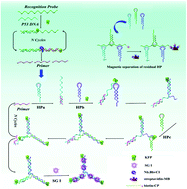
Developing a sensitive, low-cost and general sensing platform for the analysis of a DNA biomarker and its mutation is important for early cancer screening. In our work, the tumor suppressor gene–p53 DNA was chosen as the model DNA biomarker due to its vital role in preventing oncogene cancer-inhibiting activity through mediating cellular proliferation and apoptosis. Compared with tumor biopsy, the quantification of p53 DNA and its mutation in biofluids (such as urine) is more convenient due to its simple operation and non-invasiveness. Herein, a label-free amplified fluorescence assay has been developed for p53 DNA in urine samples through the KFP polymerase-driven double strand displacement reactions and a magnetic nanoprobe. First, the ssDNA probe (RP) was designed with antisense sequences for p53 DNA and the Nb.BbvCI endonuclease recognition site. In the presence of p53 DNA, the formed dsDNA between RP and p53 DNA served as an engaging primer to initiate the first strand displacement reaction (SDA) under the action of KFP DNA polymerase and Nb.BbvCI, generating abundant short ssDNA (primer). Subsequently, the resulting primers will initiate the downstream SDA through the primer–hairpin DNA (HPa) binding, opening up, and extension of HPb and HPc under the action of KFP DNA polymerase. In the process of this final DNA polymerization reaction, the primer hybridized on HPa is released and goes on to initiate another round, forming plenty of duplex Y-shaped DNA. With the integration of SYBR Green I (SG I) into these duplex DNA, the amplified label-free fluorescence detection platform for p53 DNA can be achieved. Moreover, a biotin modified nanoprobe (bio-CP) was used to capture the superfluous HP. By performing the separation function, the binding of superfluous HP and SG could be avoided and a low background can be acquired. Benefiting from the abundant SG intercalation sites of Y-shaped DNA and low background signals, this method showed excellent sensitivity with a detection limit of 0.012 nM, and the p53 DNA in urine samples was evaluated, offering a powerful tool for biomedical research and clinical diagnosis.


 Please wait while we load your content...
Please wait while we load your content...