Correlative surface imaging reveals chemical signatures for bacterial hotspots on plant roots
Abstract
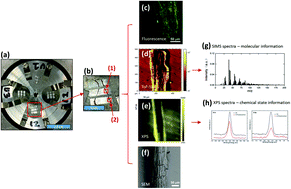
The rhizosphere is arguably the most complex microbial habitat on Earth, comprising an integrated network of plant roots, soil and a highly diverse microbial community (the rhizosphere microbiome). Understanding, predicting and controlling plant-microbe interactions in the rhizosphere will allow us to harness the plant microbiome as a means to increase or restore plant ecosystem productivity, improve plant responses to a wide range of environmental perturbations, and mitigate the effects of climate change by designing ecosystems for long-term soil carbon storage. To this end, it is imperative to develop new molecular approaches with high spatial resolution to capture interactions at the plant-microbe, microbe-microbe, and plant-plant interfaces. In this work, we designed an imaging sample holder that allows integrated surface imaging tools to map the same locations of a plant root-microbe interface with submicron lateral resolutions, providing novel in vivo analysis of root-microbe interactions. Specifically, confocal fluorescence microscopy, time-of-flight secondary ion mass spectrometry (ToF-SIMS), X-ray photoelectron spectroscopy (XPS), and scanning electron microscopy (SEM) were used for the first time for the correlative imaging of the Brachypodium distachyon root and its interaction with Pseudomonas SW25, a typical plant growth-promoting soil bacterium. Imaging data suggest that the root surface is inhomogeneous and that the interaction between Pseudomonas and Brachypodium roots was confined to only a few spots along the sampled root segments and that the bacterial attachment spots were enriched in Na- and S-related and high-mass organic species. We conclude that the attachment of the Pseudomonas cells to the root surface is outcompeted by strong root-soil mineral interactions but facilitated by the formation of extracellular polymeric substances (EPS).


 Please wait while we load your content...
Please wait while we load your content...