Novel method to identify group-specific non-catalytic pockets of human kinome for drug design†
Abstract
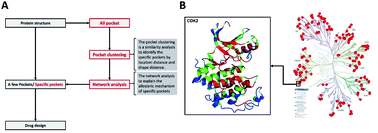
Kinase proteins have been intensively investigated as drug targets for decades because of their crucial involvement in many biological pathways. Most kinase drugs target the catalytic ATP pocket, which is highly conserved across the kinome, and as such often leads to potential side effects. It is thus highly desirable to develop non-ATP-competitive drugs that inhibit kinase activity via allosteric interactions. However, to elucidate the complex allosteric mechanism, it is essential to build a novel method to characterize a comprehensive non-catalytic pocket for the structurally well-covered human kinome. In this work, we developed a hybrid approach of sequence, structure and network analysis on 168 representative kinases to identify group-specific non-catalytic pockets. The geometric analysis was performed to cluster these pockets and to identify group-specific non-catalytic pockets based on their shape and location characteristics. Subsequent sequence evolutionary analysis reveals the crucial residues of each pocket that will likely interact with inhibitors binding to the pocket. These residues thus serve as potential biomarkers of each pocket for inhibitor design. Moreover, the residue–residue interaction network analysis was performed to elucidate the complex allosteric mechanism of these non-catalytic pockets. The final list of 14 group-specific non-catalytic pockets and their characterized structural, sequence and network features can be an enabling dataset for drug design effort at the human kinome level. The developed hybrid approach is able to identify group-specific non-catalytic pockets and will benefit the research related to human kinome drug design.


 Please wait while we load your content...
Please wait while we load your content...