Single-molecule dynamic DNA junctions for engineering robust molecular switches†
Abstract
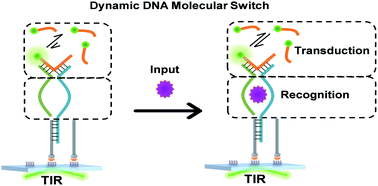
DNA molecular switches have emerged as a versatile and highly programmable toolbox and are extensively used in sensing, diagnosis, and therapeutics. Toehold mediated strand displacement serves as the core reaction for signal transduction and amplification. However, the severe leakage of this reaction limits the development of scalable and robust circuits. We engineered single-molecule dynamic DNA junctions for developing ‘zero-leakage’ molecular switches which are responsive to various inputs. Input binding enhances dynamic junctions' stability allowing for the transient binding of fluorescent probes as the output signal. Unlike the conventional intensity-based output, this molecular switch provides remarkably distinguishable kinetics-based outputs permitting ruling out leakage signals at the single-molecule level. The inputs are detected with significant sensitivity without using any amplification step. It is also revealed that the output signal is sensitive to the binding affinity of inputs and their recognition elements making the molecular switch a potential affinity meter. Considering these features, we anticipate that it would find broad applications in large-scale DNA circuits, responsive materials, and biomolecule interaction study.
- This article is part of the themed collection: 2019 Chemical Science HOT Article Collection


 Please wait while we load your content...
Please wait while we load your content...