Bisulfite-free, single base-resolution analysis of 5-hydroxymethylcytosine in genomic DNA by chemical-mediated mismatch†
Abstract
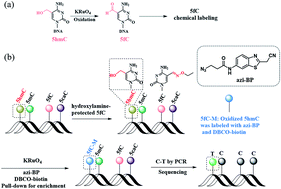
5-Hydroxymethylcytosine (5hmC) is known as one of the vital players in nuclear reprogramming and the process of active DNA demethylation. Although the development of whole-genome sequencing methods for modified cytosine bases has burgeoned, the easily operated gene-specific loci detection of 5hmC has rarely been reported. Herein, we present a single-base resolution approach, i.e., chemical-assisted mismatch sequencing (CAM-Seq), which, when combined with traditional oxidation and chemical labeling mediation, can be used for mapping 5hmC at base resolution. We employ chemical oxidation to transform 5hmC to 5-formylcytosine (5fC), followed by chemical labeling to induce C-to-T base changes owing to the fact that the loss of the exocyclic 4-amino group of labeled 5fC leads to C to T conversion and subsequent pairing with adenosine (A) in PCR. The feasibility of CAM-Seq is demonstrated in different synthetic oligonucleotide models as well as in part of the genome of 5hmC-rich mouse embryonic stem cells (mESCs). Moreover, the gene fragment containing 5hmC can be easily biotinylated after oxidation, showing high enrichment efficiency. Our method has the potential capability to map 5hmC in genomic DNA and thus will contribute to promoting the understanding of the epigenetic modification of 5hmC.


 Please wait while we load your content...
Please wait while we load your content...