Super-resolution microscopy can identify specific protein distribution patterns in platelets incubated with cancer cells†
Abstract
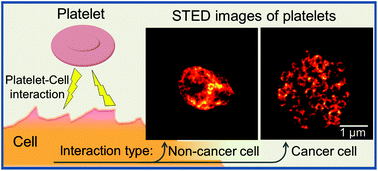
Protein contents in platelets are frequently changed upon tumor development and metastasis. However, how cancer cells can influence protein-selective redistribution and release within platelets, thereby promoting tumor development, remains largely elusive. With fluorescence-based super-resolution stimulated emission depletion (STED) imaging we reveal how specific proteins, implicated in tumor progression and metastasis, re-distribute within platelets, when subject to soluble activators (thrombin, adenosine diphosphate and thromboxane A2), and when incubated with cancer (MCF-7, MDA-MB-231, EFO21) or non-cancer cells (184A1, MCF10A). Upon cancer cell incubation, the cell-adhesion protein P-selectin was found to re-distribute into circular nano-structures, consistent with accumulation into the membrane of protein-storing alpha-granules within the platelets. These changes were to a significantly lesser extent, if at all, found in platelets incubated with normal cells, or in platelets subject to soluble platelet activators. From these patterns, we developed a classification procedure, whereby platelets exposed to cancer cells, to non-cancer cells, soluble activators, as well as non-activated platelets all could be identified in an automatic, objective manner. We demonstrate that STED imaging, in contrast to electron and confocal microscopy, has the necessary spatial resolution and labelling efficiency to identify protein distribution patterns in platelets and can resolve how they specifically change upon different activations. Combined with image analyses of specific protein distribution patterns within the platelets, STED imaging can thus have a role in future platelet-based cancer diagnostics and therapeutic monitoring. The presented approach can also bring further clarity into fundamental mechanisms for cancer cell–platelet interactions, and into non-contact cell-to-cell interactions in general.


 Please wait while we load your content...
Please wait while we load your content...