A method for SNP detection using MoS2@AuNPs and SYBR Green I in combination with enzyme digestion
Abstract
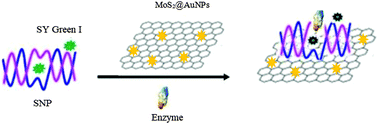
Single-nucleotide polymorphisms (SNPs) in a gene sequence are markers for a variety of human diseases. SNP detection methods are time-consuming, need laboratory-scale settings, and are expensive. A simple, rapid and label-free method for detecting SNPs was developed in this study. SG (SYBR Green I), DNA sequences, and molybdenum disulfide–gold nanoparticles (MoS2@AuNPs) were used in this sensing system for a proof-of-principle study. DNA was immobilized on the surface of MoS2@AuNPs in order to remove the nonspecific probe displacement and false-positive signals using 10% Tween 80. MoS2@AuNPs quenched the fluorescence from SG intercalated into the DNA duplex. The SG fluorescence signals from the matched and mismatched DNA duplex were significantly different after the addition of MoS2. HapII can recognize the enzyme digestion site in the DNA sequence and further improve the effect of SNP detection. Therefore, this method can clearly distinguish SNPs in DNA sequences.


 Please wait while we load your content...
Please wait while we load your content...