A comprehensive profile of the tilapia (Oreochromis niloticus) circular RNA and circRNA–miRNA network in the pathogenesis of meningoencephalitis of teleosts†
Abstract
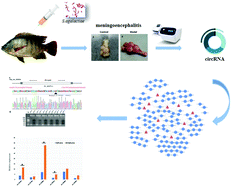
The pathogenesis of tilapia meningoencephalitis is still unclear, where the involvement of circRNA is considered for its active role as a “miRNA sponge”. Therefore, we aimed to investigate the profile of circRNA in tilapia meningoencephalitis further by constructing the circRNA–miRNA network for in-depth mechanism exploration. Briefly, a nile tilapia model of meningitis was established by injecting Streptococcus agalactiae (1.0 × 107 cfu per mL) and we evaluated the infected tilapia brain for the expression profile of circRNAs, their potential functions and their correlation with genes involved in inflammatory pathways. A total of 11 263 circRNAs were identified from RNA sequencing (RNA-seq) data in nile tilapia (Oreochromis niloticus), a commercially important fish in China and East Asia. GO and KEGG analyses revealed that the biological functions of genes hosting the circRNAs were enriched in the progression of metabolism and binding. Notably, we found that 99% circRNAs in tilapia had abundant miRNA-binding sites and a total of 2136 of the identified circRNAs had two to six miRNA-binding sites. Six circRNAs were validated by qRT-PCR and the final circRNA–miRNA network was constructed. This is the first report of comprehensive identification of O. niloticus circRNAs being differentially regulated in the brain in normal conditions relating to S. agalactiae infection. This work will shed novel light on gene expression mechanisms under disease conditions and may identify circRNAs as new biomarkers for meningoencephalitis and neurodegenerative disorders.


 Please wait while we load your content...
Please wait while we load your content...