Cyclic dipeptide peroxosolvates: first direct evidence for hydrogen bonding between hydrogen peroxide and a peptide backbone†
Abstract
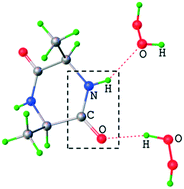
Herein, peroxosolvates of the cyclic dipeptides disarcosine (C6H10N2O2·H2O2), dialanine (C6H10N2O2·2(H2O2)), diglycine (C4H6N2O2·2(H2O2) and C4H6N2O2·1.786(H2O2)·0.214(H2O)) were synthesized and studied by single crystal X-ray analysis and periodic DFT calculations. The hydrogen bonding networks of cyclic diglycine peroxosolvate and isomorphous hydrate were characterized using Bader analysis of periodic electron density and empirical correlations between enthalpy and metric and spectroscopic parameters of the H-bond. The obtained data for the peroxosolvate of cyclic dipeptides provided a molecular-level insight into the non-redox interaction of hydrogen peroxide with the peptide backbone in aqueous systems. In all structures, H2O2 acts as a proton donor in two hydrogen bonds. The carbonyl oxygen atom of the peptide (–CO–NH–) forms a relatively stronger hydrogen bond with the hydrogen peroxide molecule (32–34 kJ mol−1) as compared to that with water in the isomorphous hydrate of cyclic diglycine (26–28 kJ mol−1). The hydrogen atom of the peptide group (–CO–NH–) can form a moderate H-bond with hydrogen peroxide (25 kJ mol−1). The total hydrogen bonding energy of the peroxosolvate is higher than that of the corresponding hydrate (91 vs. 82 kJ mol−1). The stronger hydrogen bonding in peroxosolvates as compared to that in the hydrate and the fact that the intermolecular H-bonds C(![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O)NH⋯O
O)NH⋯O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CNH between cyclic dipeptides are not found in peroxosolvates, despite being prevalent in the structure of nonsolvated dipeptides, imply that hydrogen peroxide can interfere with the secondary structure of proteins via a mechanism that can lead to reversible inhibition or signaling in living systems.
CNH between cyclic dipeptides are not found in peroxosolvates, despite being prevalent in the structure of nonsolvated dipeptides, imply that hydrogen peroxide can interfere with the secondary structure of proteins via a mechanism that can lead to reversible inhibition or signaling in living systems.


 Please wait while we load your content...
Please wait while we load your content...