Rapid identification of specific DNA aptamers precisely targeting CD33 positive leukemia cells through a paired cell-based approach†
Abstract
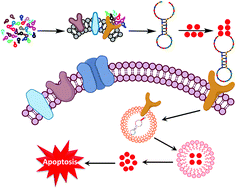
Aptamers are short single-stranded DNA or RNA molecules, which have recently been developed for potential broad applications such as clinical therapeutics, diagnosis and tumor-targeted drug delivery. However, the selection of specific aptamers is often unsatisfactory using the classical protein or cell-based SELEX. Herein, we modified the paired cell line approach to identify aptamers targeting leukemia cells expressing the CD33 antigen. Our strategy artfully used the same cells for negative (HEK293T cells) and positive (CD33 transfected-HEK293T cells) aptamer selections, and the negative selections were performed adequately before the positive selection to remove unspecific sequences. The advantages of this strategy are that it is fast and accurate, where only a few rounds of selection together with PCR amplifications are sufficient to obtain high binding affinity antigen-targeted aptamers. By using our modified approach, we successfully obtained the CD33-targeting aptamer S30, which could highly recognize the C2 domain of the CD33 antigen in vitro and in vivo. Moreover, the optimized aptamer S30-T1 (i.e., core region of S30) was conjugated with doxorubicin (Dox) to synthesize S30-T1-Dox conjugates, which could specifically inhibit CD33 positive acute myeloid leukemia HL-60 cell proliferation by arresting the cell cycle at the G2 phase. Thus, our modified approach can rapidly screen reliable, stable and high binding affinity aptamers for precise cancer treatment.


 Please wait while we load your content...
Please wait while we load your content...