Single molecule analysis of structural fluctuations in DNA nanostructures†
Abstract
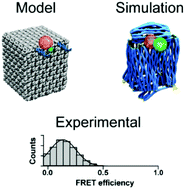
DNA origami is an excellent tool for building complex artificial nanostructures. Functionalization of these structures provides the possibility of precise organization of matter at the nanoscale. In practice, efforts in this endeavour can be impeded by electrostatic repulsion or other dynamics at the molecular scale, resulting in uncompliant local structures. Using single molecule FRET microscopy combined with coarse-grained Brownian dynamics simulations, we investigated here the local structure around the lid of a DNA origami box, which can be opened by specific DNA keys. We found that FRET signals for the closed box depend on buffer ion concentrations and small changes to the DNA structure design. Simulations provided a view of the global and local structure and showed that the distance between the box wall and lid undergoes fluctuations. These results provide methods to vizualise and improve the local structure of three-dimensional DNA origami assemblies and offer guidance for exercising control over placement of chemical groups and ligands.


 Please wait while we load your content...
Please wait while we load your content...
