Relationship between solid state structure and solution stability of copper(ii)–hydroxypyridinecarboxylate complexes†
Abstract
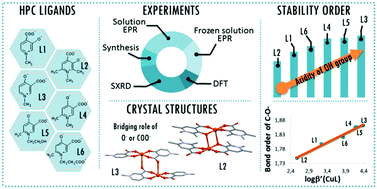
The complementary solid state/solution studies of the systematic series of bioactive ligands 3-hydroxy-1-methyl-4-pyridinecarboxylate (L1), 3-hydroxy-1,2,6-trimethyl-4-pyridinecarboxylate (L2), 4-hydroxy-1-methyl-3-pyridinecarboxylate (L3), 4-hydroxy-1,6-dimethyl-3-pyridinecarboxylate (L4), 4-hydroxy-1-(2-hydroxyethyl)-6-methyl-3-pyridinecarboxylate (L5) and 4-hydroxy-1-(2-carboxyethyl)-6-methyl-3-pyridinecarboxylate (L6) with copper(II) have been performed in order to design efficient chelating drugs for the treatment of metal overloading conditions. Single crystals of [Cu(L1)2(H2O)]·3H2O (1) (monomer) with axial water coordination, [Cu2(L2)4]·6H2O (2) and [Cu2(L3)4]·4H2O (3) (cyclic dimers), where pyridinolato and carboxylato oxygens, respectively, act as linkers between adjacent copper complexes, [Cu(L4)2]n·3H2O (4) (1D polymer) and [Cu3(L5)6]·18H2O (5) (trimer), constructed using two square-pyramidal and one elongated octahedral Cu(II) complexes have been determined by SXRD. The bidentate coordination mode of the ligands has been found preferentially with cis arrangements in 1 and 2 and trans arrangements in 3–5. The solution speciation and complex stability of aqueous solutions have been studied by pH-dependent electron paramagnetic resonance spectroscopy resulting in the detection of solely monomeric [CuL]+ and [CuL2] complexes. The stability order obtained for the [CuL]+ complexes could be correlated with the deprotonation constants of their hydroxyl group (log βLH) reflecting that the higher acidity increases the complex stability in the order L2 < L1 ≈ L6 < L4 ≈ L5 < L3. This stability order elucidates the different axial linkers in the cyclic dimers 2 and 3. DFT quantum-chemical calculations support the effect of the electron distribution on the established stability order.


 Please wait while we load your content...
Please wait while we load your content...