Simulation-guided engineering of an enzyme-powered three dimensional DNA nanomachine for discriminating single nucleotide variants†
Abstract
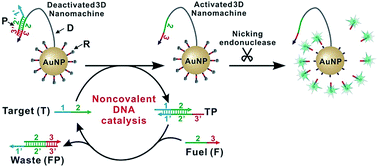
Single nucleotide variants (SNVs) are important both clinically and biologically because of their profound biological consequences. Herein, we engineered a nicking endonuclease-powered three dimensional (3D) DNA nanomachine for discriminating SNVs with high sensitivity and specificity. Particularly, we performed a simulation-guided tuning of sequence designs to achieve the optimal trade-off between device efficiency and specificity. We also introduced an auxiliary probe, a molecular fuel capable of tuning the device in solution via noncovalent catalysis. Collectively, our device produced discrimination factors comparable with commonly used molecular probes but improved the assay sensitivity by ∼100 times. Our results also demonstrate that rationally designed DNA probes through computer simulation can be used to quantitatively improve the design and operation of complexed molecular devices and sensors.
- This article is part of the themed collection: Most popular 2018-2019 nanoscience articles


 Please wait while we load your content...
Please wait while we load your content...