Electron transfer ferredoxins with unusual cluster binding motifs support secondary metabolism in many bacteria†
Abstract
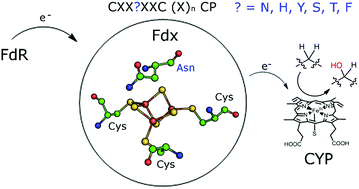
The proteins responsible for controlling electron transfer in bacterial secondary metabolism are not always known or characterised. Here we demonstrate that many bacteria contain a set of unfamiliar ferredoxin encoding genes which are associated with those of cytochrome P450 (CYP) monooxygenases and as such are involved in anabolic and catabolic metabolism. The model organism Mycobacterium marinum M contains eleven of these genes which encode [3Fe–4S] or [4Fe–4S] single cluster containing ferredoxins but which have unusual iron–sulfur cluster binding motif sequences, CXX?XXC(X)nCP, where ‘?’ indicates a variable amino acid residue. Rather than a cysteine residue, which is highly conserved in [4Fe–4S] clusters, or alanine or glycine residues, which are common in [3Fe–4S] ferredoxins, these genes encode at this position histidine, asparagine, tyrosine, serine, threonine or phenylalanine. We have purified, characterised and reconstituted the activity of several of these CYP/electron transfer partner systems and show that all those examined contain a [3Fe–4S] cluster. Furthermore, the ferredoxin used and the identity of the variable motif residue in these proteins affects the functionality of the monooxygenase system and has a significant influence on the redox properties of the ferredoxins. Similar ferredoxin encoding genes were identified across Mycobacterium species, including in the pathogenic M. tuberculosis and M. ulcerans, as well as in a wide range of other bacteria such as Rhodococcus and Streptomyces. In the majority of instances these are associated with CYP genes. These ferredoxin systems are important in controlling electron transfer across bacterial secondary metabolite production processes which include antibiotic and pigment formation among others.


 Please wait while we load your content...
Please wait while we load your content...