Unraveling the 4n − 1 rule for DNA i-motif stability: base pairs vs. loop lengths†
Abstract
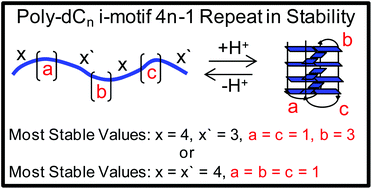
Previously our laboratory identified that poly-2′-deoxycytidine (dCn) strands of DNA with lengths greater than 12 nucleotides could adopt i-motif folds, while the pH-dependent stabilities follow a 4n − 1 repeat pattern with respect to chain length (J. Am. Chem. Soc., 2017, 139, 4682–4689). Herein, model i-motif folds in which loop configurations were forced by judiciously mutating dC to non-dC nucleotides allowed a structural model to be proposed to address this phenomenon. The model was developed by systematically studying two i-motifs with either an even or odd number of d(C·C)+ hemiprotonated base pairs in the core. First, a trend in the pH-dependent stability vs. loop nucleotide identity was observed: dC > dT ∼ dU ≫ dA ∼ dG. Next, loops comprised of dT nucleotides in the two different core base pair configurations were studied while systematically changing the loop lengths. We found that an i-motif with an even number of base pairs in the core with a single nucleotide in each of the three loops was the most stable, as well as an i-motif with an odd number of core base pairs having one nucleotide in the two exterior loops and three nucleotides in the central loop. A systematic increase in the central loop from 1–4 nucleotides for an odd number of base pairs in the i-motif core reproduced the 4n − 1 repeat pattern observed in the poly-dCn strands. Additional loop configurations were studied to further support the model. The results are discussed with respect to their biological relevance.
- This article is part of the themed collections: Chemical Biology in OBC and Celebrating excellence in research: women of organic chemistry


 Please wait while we load your content...
Please wait while we load your content...