Binding of aluminium/cacodylate complexes with DNA and RNA. Experimental and “in silico”study†
Abstract
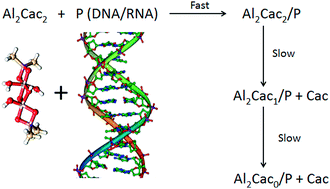
The interaction of RNA and DNA with two aluminium-containing complexes has been studied at 25 °C using 1H-NMR, UV and CD spectroscopies, viscosity measurements and DFT simulations. The complex identified at pH = 7, Al13Cacx, can interact with polynucleotides by external binding. Even though the affinity of the aluminium complex for RNA and DNA is similar, the conformational changes induced upon interaction are greater with RNA. In contrast, at pH = 5 the complex present in solution is Al2Cac2, where the aluminium atoms have different configurations, such as octahedral (OH) and distorted square planar (DSP). Al2Cac2 is prone to interact covalently with both P (RNA or DNA) to yield Al2Cac2/P complexes. These complexes undergo slow decomposition in two consecutive steps to give Al2Cac1/P after losing the cacodylate moiety belonging to the Al(OH) and Al2Cac0/P when the second cacodylate residue of the Al(DSP) is lost. These steps are (roughly) twice as fast with DNA than with RNA and for both systems the release of the first cacodylate molecule is about 10 times faster than the second. The cacodylate present in the buffer acts not only as a complexing agent of Al but also as an active vehicle that leads the metal to its biological target; once the target is reached, the two cacodylate molecules are released, leaving Al covalently linked to the N7 of guanine and adenine. These features reveal the great complexity of aluminium interaction with biological systems, an important outcome due to the implications of this metal in human health.
- This article is part of the themed collection: Equilibrium Solution Coordination Chemistry


 Please wait while we load your content...
Please wait while we load your content...