Formation of arrays of planar, murine, intestinal crypts possessing a stem/proliferative cell compartment and differentiated cell zone†
Abstract
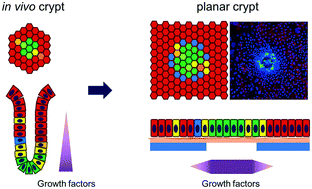
A simple, in vitro intestinal model recapitulating key aspects of crypt architecture and physiology would facilitate our understanding the impact of drugs, foods and microbial metabolites on the intestine. To address the limitations of previously reported intestinal in vitro platforms, we developed a planar crypt array that replicated the spatial segregation and physiologic responses of primary mouse intestinal epithelial cells in the large intestine. Collagen was coated across an impermeable film possessing an array of microholes creating two regions of distinct stiffness and porosity (above and outside the microholes). Primary mouse colon epithelial cells formed a continuous monolayer across the array with a proliferative cell zone above the microholes and a nonproliferative or differentiated cell region distant from the microholes. Formation of a chemical gradient of growth factors across the array yielded a more complete or in vivo-like cell segregation of proliferative and differentiated cells with cell migration outward from the proliferative cell zone into the differentiated zone to replace apoptotic dying cells much as occurs in vivo. Short chain fatty acids (microbial metabolites) applied to the luminal surface of the crypt array significantly impacted the proliferation and differentiation of the cells replicating the known in vivo effects of these fatty acids. Importantly this planar crypt array was readily fabricated and maintained, easily imaged with properties quantified by microscopy, and compatible with reagent addition to either the luminal or basal fluid reservoirs. The ability to observe simultaneously stem/proliferative and differentiated cell behavior and movement between these two compartments in response to drugs, toxins, inflammatory mediators or microbial metabolites will be of widespread utility.
- This article is part of the themed collection: Organ-, body- and disease-on-a-chip systems


 Please wait while we load your content...
Please wait while we load your content...