A new photoelectrochemical biosensor for ultrasensitive determination of nucleic acids based on a three-stage cascade signal amplification strategy†
Abstract
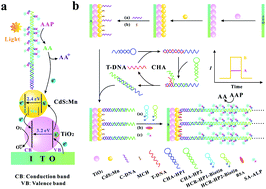
The sensitive and specific determination of nucleic acids is very important in clinical diagnosis and biological studies. In this work, an ultrasensitive photoelectrochemical (PEC) biosensor has been developed for DNA detection based on a “signal-on” sensing strategy and a three-stage cascade signal amplification method (catalytic hairpin assembly (CHA), hybridization chain reaction (HCR) and alkaline phosphatase (ALP)-triggered in situ generation of ascorbic acid (AA)). Here, CHA hairpin 1 (CHA-HP1) is opened by the target DNA (T-DNA) owing to the hybridization between T-DNA and CHA-HP1, and then the opened CHA-HP1 hybridizes with CHA hairpin 2 (CHA-HP2) to displace the T-DNA, generating a CHA-HP1/CHA-HP2 complex. The displaced T-DNA triggers the next cycle of CHA, resulting in the generation of numerous CHA-HP1/CHA-HP2 complexes. Subsequently, one end of the CHA-HP1/CHA-HP2 complex hybridizes with the capture DNA immobilized on the indium tin oxide/TiO2/CdS : Mn electrode. After the introduction of dual-biotin labeled HCR hairpin 1 (HCR-HP1-Bio) and dual-biotin labeled HCR hairpin 2 (HCR-HP2-Bio), the other end of the CHA-HP1/CHA-HP2 complex opens HCR-HP1-Bio. The opened HCR-HP1-Bio triggers the HCR reaction between HCR-HP1-Bio and HCR-HP2-Bio, leading to the formation of long nicked duplex DNA structures. The dual-biotin modified HCR-hairpins can anchor more streptavidin–ALP to catalyze 2-phospho-L-ascorbic acid trisodium salt to yield more AA, leading to a larger PEC response. The proposed PEC biosensor shows superior analytical performance for T-DNA detection with a linear response ranging from 0.1 fM to 100 pM and a detection limit of 0.052 fM, and may provide a powerful biosensing platform for bioanalysis and early disease diagnosis.
- This article is part of the themed collections: Analyst Recent HOT articles, Analytical Nanoscience and Recent analytical chemistry science from China


 Please wait while we load your content...
Please wait while we load your content...