A novel restriction endonuclease GlaI for rapid and highly sensitive detection of DNA methylation coupled with isothermal exponential amplification reaction†
Abstract
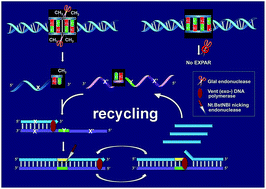
Sensitive and accurate detection of site-specific DNA methylation is of critical significance for early diagnosis of human diseases, especially cancers. Herein, for the first time we employ a novel methylation-dependent restriction endonuclease GlaI to detect site-specific DNA methylation in a highly specific and sensitive way by coupling with isothermal exponential amplification reaction (EXPAR). GlaI can only cut the methylated target site with excellent selectivity but leave the unmethylated DNA intact. Then the newly exposed end fragments of methylated DNA can trigger EXPAR for highly efficient signal amplification while the intact unmethylated DNA will not initiate EXPAR at all. As such, only the methylated DNA is quantitatively and faithfully reflected by the real-time fluorescence signal of the GlaI–EXPAR system, and the potential false positive interference from unmethylated DNA can be effectively eliminated. Therefore, by integrating the unique features of GlaI for highly specific methylation discrimination and EXPAR for rapid and powerful signal amplification, the elegant GlaI–EXPAR assay allows the direct quantification of methylated DNA with ultrahigh sensitivity and accuracy. The detection limit of methylated DNA target has been pushed down to the aM level and the whole detection process of GlaI–EXPAR can be accomplished within a short time of 2 h. More importantly, ultrahigh specificity is achieved and as low as 0.01% methylated DNA can be clearly identified in the presence of a large excess of unmethylated DNA. This GlaI–EXPAR is also demonstrated to be capable of determining site-specific DNA methylations in real genomic DNA samples. Sharing the distinct advantages of ultrahigh sensitivity, outstanding specificity and facile operation, this new GlaI–EXPAR strategy may provide a robust and reliable platform for the detection of site-specific DNA methylations with low abundances.


 Please wait while we load your content...
Please wait while we load your content...