Defining the current scope and limitations of dual noncanonical amino acid mutagenesis in mammalian cells†
Abstract
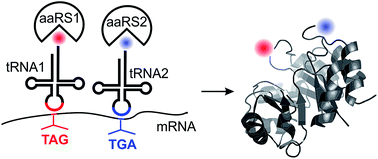
The ability to site-specifically incorporate two distinct noncanonical amino acids (ncAAs) into the proteome of a mammalian cell with high fidelity and efficiency will have many enabling applications. It would require the use of two different engineered aminoacyl-tRNA synthetase (aaRS)/tRNA pairs, each suppressing a distinct nonsense codon, and which cross-react neither with each other, nor with their counterparts from the host cell. Three different aaRS/tRNA pairs have been developed so far to expand the genetic code of mammalian cells, which can be potentially combined in three unique ways to drive site-specific incorporation of two distinct ncAAs. To explore the suitability of using these combinations for suppressing two distinct nonsense codons with high fidelity and efficiency, here we systematically investigate: (1) how efficiently the three available aaRS/tRNA pairs suppress the three different nonsense codons, (2) preexisting cross-reactivities among these pairs that would compromise their simultaneous use, and (3) whether different nonsense-suppressor tRNAs exhibit unwanted suppression of non-cognate stop codons in mammalian cells. From these comprehensive analyses, two unique combinations of aaRS/tRNA pairs emerged as being suitable for high-fidelity dual nonsense suppression. We developed expression systems to validate the use of both combinations for the site-specific incorporation of two different ncAAs into proteins expressed in mammalian cells. Our work lays the foundation for developing powerful applications of dual-ncAA incorporation technology in mammalian cells, and highlights aspects of this nascent technology that need to be addressed to realize its full potential.


 Please wait while we load your content...
Please wait while we load your content...