Insights into the RNA binding mechanism of human L1-ORF1p: a molecular dynamics study†
Abstract
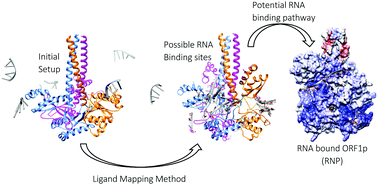
The recognition and binding of nucleic acids by ORF1p, an L1 retrotransposon protein, have not yet been clearly understood due to the lack of structural knowledge. The present study attempts to identify the probable single-stranded RNA binding pathway of trimeric ORF1p using computational methods like ligand mapping methodology combined with molecular dynamics simulations. Using the ligand mapping methodology, the possible RNA interacting sites on the surface of the trimeric ORF1p were identified. The crystal structure of the ORF1p timer and an RNA molecule of 29 nucleotide bases in length were used to generate the structure of the ORF1p complex based on information on predicted binding sites as well as the functional states of the CTD. The various complexes of ORF1p–RNA were generated using polyU, polyA and L1RNA sequences and were simulated for a period of 75 ns. The observed stable interaction pattern was used to propose the possible binding pathway. Based on the binding free energy for complex formation, both polyU and L1RNA complexes were identified as stable complexes, while the complex formed with polyA was the least stable one. Furthermore, the importance of the residues in the CC domain (Lys137 and Arg141), the RRM loop (Arg206, Arg210 and Arg211) and the CTD (Arg 261 and Arg262) of all three chains in stabilizing the wrapped RNA has been highlighted in this study. The presence of several electrostatic interactions including H-bond interactions increases the affinity towards RNA and hence plays a vital role in retaining the wrapped position of RNA around ORF1p. Altogether, this study presents one of the possible RNA binding pathways of ORF1p and clearly highlights the functional state of ORF1p visited during RNA binding.


 Please wait while we load your content...
Please wait while we load your content...