In situ mRNA isolation from a microfluidic single-cell array using an external AFM nanoprobe†
Abstract
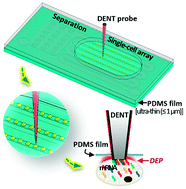
We present an in situ mRNA extraction platform to quantify marker-genes' expression levels of single target cells within high-density microfluidic trapping arrays. This platform enables single-cell transcriptomic analysis to reveal in-depth information of cellular mechanisms and population heterogeneity. Although microfluidic technology enables the automation of single-cell sorting, trapping and identification, most developed microfluidic devices are closed off and prevent single-cell access by external analytical equipment. Besides, cell lysing is usually required for mRNA extraction. In our platform, cells are trapped individually in a microwell array sealed by a 1 μm-thick polydimethylsiloxane (PDMS) membrane, and a modified atomic force microscopy (AFM) probe—a dielectrophoretic nanotweezer (DENT)—penetrates through the membrane and extracts mRNA molecules from a single cell by dielectrophoresis. The single-cellular expression levels of 3 housekeeping genes from HeLa cells were analyzed quantitatively based on the quantification of the extracted mRNAs, and the probed cells remained viable when the applied alternating-current (AC) voltage was lower than 1.5 Vpp during mRNA probing. We also performed in situ mRNA isolation from a mixture of SK-BR-3 and U937 cells, mimicking a blood sample that underwent primary enrichment of circulating tumor cells (CTCs), and evaluated various marker-genes' expressions. This integrated platform combines the non-destructive and precise-control of a single-cell mRNA probe with sealed microfluidic systems' capability of upstream sample processing and downstream multifunctional analysis to enable a versatile and powerful tool for biomedical research.
- This article is part of the themed collection: RSC papers by NanoEngineering for Medicine and Biology 2018 Speakers


 Please wait while we load your content...
Please wait while we load your content...