Extracting DNA from ocean microplastics: a method comparison study†
Abstract
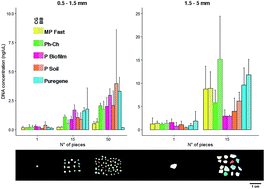
The ubiquity of plastics in oceans worldwide raises concerns about their ecological implications. Suspended microplastics (<5 mm) can be ingested by a wide range of marine organisms and may accumulate up the food web along with associated chemicals. Additionally, plastics provide a stable substrate to a wide range of organisms and, owing to their widespread dispersal, may function as vectors for harmful and invasive species. Despite the growing application of molecular techniques to study ocean microplastic colonizers, to date there is no comparative study on DNA extraction methods for ocean plastic biofilms. The present study aims to fill this gap by comparing DNA yield, amplification efficiency, costs and processing time of different DNA extraction techniques applied to oceanic microplastics. DNA was extracted with five methods (four extraction kits, and standard phenol:chloroform purification) using two mechanical lysis techniques (bead beating and cryogenic grinding with liquid nitrogen) applied to three plastic quantities (1, 15, and 50 fragments per extraction) and size classes (0.05–0.15 and 0.15–0.5 mm). All methods resulted in DNA suitable for downstream applications and were successfully amplified. Overall, the Qiagen Puregene Tissue kit yielded relatively high DNA concentrations for most sizes and amounts of plastics at relatively low costs and short processing time. This study provides a detailed evaluation of DNA extraction methods from ocean plastics, and may assist future research using molecular techniques to study ocean plastic biofilms.
- This article is part of the themed collections: Analytical Methods 2017 Most Downloaded Articles and Microplastics in the environment


 Please wait while we load your content...
Please wait while we load your content...