Local water sensing: water exchange in bacterial photosynthetic reaction centers embedded in a trehalose glass studied using multiresonance EPR†
Abstract
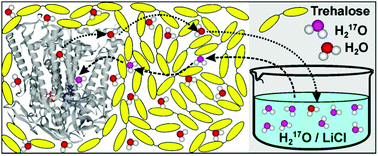
Using isotope labeled water (D2O and H217O) and pulsed W-band (94 GHz) high-field multiresonance EPR spectroscopies, such as ELDOR-detected NMR and ENDOR, the biologically important question of detection and quantification of local water in proteins is addressed. A bacterial reaction center (bRC) from Rhodobacter sphaeroides R26 embedded into a trehalose glass matrix is used as a model system. The bRC hosts the two native radical cofactor ions 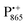 (primary electron donor) and
(primary electron donor) and 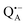 (primary electron acceptor) as well as an artificial nitroxide spin label site-specifically attached to the surface of the H-protein domain. The three paramagnetic reporter groups have distinctly different local environments. They serve as local probes to detect water molecules via magnetic interactions (electron–nuclear hyperfine and quadrupole) with either deuterons or 17O nuclei. bRCs were equilibrated in an atmosphere of different relative humidities allowing us to control precisely the hydration levels of the protein. We show that by using oxygen-17 labeled water quantitative conclusions can be made in contrast to using D2O which suffers from proton–deuterium exchange processes in the protein. From the experiments we also conclude that dry trehalose operates as an anhydrobiotic protein stabilizer in line with the “anchorage hypothesis” of bio-protection. It predicts selective changes in the first solvation shell of the protein upon trehalose–matrix dehydration with subsequent changes in the hydrogen-bonding network. Changes in hydrogen-bonding patterns usually have an impact on the global function of a biological system.
(primary electron acceptor) as well as an artificial nitroxide spin label site-specifically attached to the surface of the H-protein domain. The three paramagnetic reporter groups have distinctly different local environments. They serve as local probes to detect water molecules via magnetic interactions (electron–nuclear hyperfine and quadrupole) with either deuterons or 17O nuclei. bRCs were equilibrated in an atmosphere of different relative humidities allowing us to control precisely the hydration levels of the protein. We show that by using oxygen-17 labeled water quantitative conclusions can be made in contrast to using D2O which suffers from proton–deuterium exchange processes in the protein. From the experiments we also conclude that dry trehalose operates as an anhydrobiotic protein stabilizer in line with the “anchorage hypothesis” of bio-protection. It predicts selective changes in the first solvation shell of the protein upon trehalose–matrix dehydration with subsequent changes in the hydrogen-bonding network. Changes in hydrogen-bonding patterns usually have an impact on the global function of a biological system.


 Please wait while we load your content...
Please wait while we load your content...