Groove modification of siRNA duplexes to elucidate siRNA–protein interactions using 7-bromo-7-deazaadenosine and 3-bromo-3-deazaadenosine as chemical probes†
Abstract
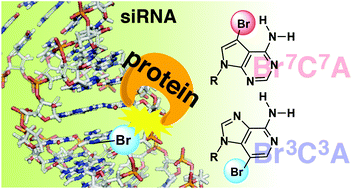
Elucidation of dynamic interactions between RNA and proteins is essential for understanding the biological processes regulated by RNA, such as RNA interference (RNAi). In this study, the logical chemical probes, comprising 7-bromo-7-deazaadenosine (Br7C7A) and 3-bromo-3-deazaadenosine (Br3C3A), to investigate small interfering RNA (siRNA)–RNAi related protein interactions, were developed. The bromo substituents of Br7C7A and Br3C3A are expected to be located in the major and the minor grooves, respectively, and to act as a steric hindrance in each groove when these chemical probes are incorporated into siRNAs. A comprehensive investigation using siRNAs containing these chemical probes revealed that (i) Br3C3A(s) at the 5′-end of the passenger strand enhanced their RNAi activity, and (ii) the direction of RISC assembly is determined by the interaction between Argonaute2, which is the main component of RISC, and siRNA in the minor groove near the 5′-end of the passenger strand. Utilization of these chemical probes enables the investigation of the dynamic interactions between RNA and proteins.

 Please wait while we load your content...
Please wait while we load your content...