New pyridazine-based binuclear nickel(ii), copper(ii) and zinc(ii) complexes as prospective anticancer agents†
Abstract
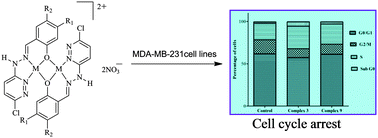
A new class of pyridazine-based binuclear nickel(II), copper(II) and zinc(II) complexes of the type [M2(L1–3)2](NO3)2 (1–9) with tridentate Schiff base ligands 3-chloro-6-(salicylidenehydrazinyl)pyridazine (HL1), 3-chloro-6-(5-nitrosalicylidenehydrazinyl)pyridazine (HL2) and 3-chloro-6-(4-diethylaminosalicylidenehydrazinyl)pyridazine (HL3) was synthesized and characterized. The molecular structure of the ligand HL3 was determined by the single crystal XRD method. The geometry optimization and HOMO–LUMO energy level calculations were carried out using DFT studies. The electrochemical studies of nickel(II) (1–3) and copper(II) (4–6) complexes exhibit two irreversible one-electron reduction waves at 1Epc = −0.323 to −0.463 V and 2Epc = −0.634 to −0.790 V in the cathodic potential region. The nickel(II) complexes (1–3) exhibit two irreversible one-electron oxidation waves at 1Epa = 1.009 to 1.025 V and 2Epa = 1.140 to 1.153 V in the anodic potential region. The spectroscopic data indicate that the ligands behave as a monoanionic tridentate ligand through the deprotonated phenolic oxygen and nitrogen atoms of the azomethine group and the pyridazine ring. The emission studies of the complexes at room temperature indicate the enhancement of fluorescence intensity than the ligands due to the chelation-enhanced fluorescence effect (CHEF) suggesting them as possible fluorescent probes of these Schiff bases for metal ions. In vitro cytotoxic activity of all the complexes was assessed against one breast cancer (MDA-MB-231) cell line and one normal cell line (L-6 myoblast). The IC50 values of complexes 3 and 9 indicate their high cytotoxicity against MDA-MB-231 cells when compared to the standard drug cisplatin suggesting that these complexes may act as potential antitumor agents. Nuclear chromatin cleavage has also been observed with AO/EB staining assay. Flow cytometry analysis of complexes 3 and 9 revealed cell cycle arrest in the S phase. Molecular docking studies were also carried out for the complexes 3 and 9 to find their binding affinity with protein EGFR kinase.

 Please wait while we load your content...
Please wait while we load your content...