High-throughput proteomics and the fight against pathogens
Abstract
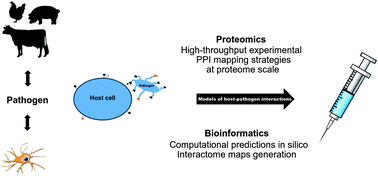
Pathogens pose a major threat to human and animal welfare. Understanding the interspecies host–pathogen protein–protein interactions could lead to the development of novel strategies to combat infectious diseases through the rapid development of new therapeutics. The first step in understanding the host–pathogen crosstalk is to identify interacting proteins in order to define crucial hot-spots in the host–pathogen interactome, such as the proposed pharmaceutical targets by means of high-throughput proteomic methodologies. In order to obtain holistic insight into the inter- and intra-species bimolecular interactions, apart from the proteomic approach, sophisticated in silico modeling is used to correlate the obtained large data sets with other omics data and clinical outcomes. Since the main focus in this area has been directed towards human medicine, it is time to extrapolate the existing expertise to a new emerging field: the ‘systems veterinary medicine’. Therefore, this review addresses high-throughput mass spectrometry-based technology for monitoring protein–protein interactions in vitro and in vivo and discusses pathogen cultivation, model host cells and available bioinformatic tools employed in vaccine development.
- This article is part of the themed collection: Omics in Animal Sciences


 Please wait while we load your content...
Please wait while we load your content...