Engineering of a fungal nitrilase for improving catalytic activity and reducing by-product formation in the absence of structural information
Abstract
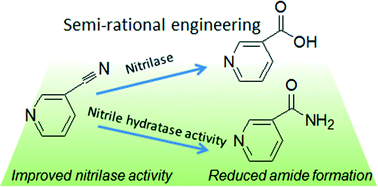
Most available methods for modifying the catalytic properties of enzymes are costly and time-consuming, as they rely on the information of enzyme crystal structure or require handling large amounts of mutants. This study employs sequence analysis and saturation mutagenesis to improve the catalytic activity and reduce the by-product formation of fungal nitrilase in the absence of structural information. Site-saturation mutagenesis of isoleucine 128 and asparagine 161 in the fungal nitrilase from Gibberella intermedia was performed and mutants I128L and N161Q showed higher catalytic activity toward 3-cyanopyridine and weaker amide forming ability than the wild-type. Moreover, the activity of double mutant I128L–N161Q was improved by 100% and the amount of amide formed was reduced to only one third of that of the wild-type. The stability of the mutants was significantly enhanced at 30 and 40 °C. The catalytic efficiency of the mutant enzymes was substantially improved. In this study, we successfully applied a novel approach that required no structural information and minimal workload of mutant screening for engineering of fungal nitrilase.

 Please wait while we load your content...
Please wait while we load your content...