Dinuclear nickel complexes of divergent Ni⋯Ni separation showing ancillary ligand addition and bio-macromolecular interaction†
Abstract
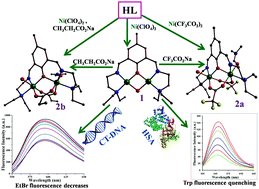
Schiff base ligand HL has been utilized to build up a new family of five [Ni2] complexes, [Ni2(μ-L)(μ-OH)](ClO4)2 [1(ClO4)2], [Ni2(μ-L)(μ-CF3CO2)2(C3H7NO)(H2O)]CF3CO2·H2O [2a(CF3CO2)·H2O], [Ni2(μ-L)(μ-CH3CH2CO2)2(C3H7NO)(H2O)]ClO4·C3H7NO [2b(ClO4)·C3H7NO], [Ni2(μ-L)(μ-CH3CO2)(μ-NCS)(NCS)(C3H7NO)] (3a) and [Ni2(μ-L)(μ-CH3CO2)(μ-N3)(N3)(C3H7NO)] (3b), in varying co-ligand environments. All the complexes have been characterized by elemental analyses, X-ray structure analysis and spectroscopic measurements. Three distinct types, dicationic, monocationic and neutral complexes, are synthesized using one pentadentate ligand HL. The NiII centers are bridged by three sets of bridging groups resulting in a wide variation in Ni⋯Ni separations from 2.897 to 3.475 Å. Doubly bridged NiII centers in the edge-shared square planar environment provide the shortest Ni⋯Ni separation in 12+. Whereas combination of other bridges from carboxylate, azide and isothiocyanate ancillary ligands results in cationic and neutral [Ni2] complexes 2+ and 3 having longer intermetallic separations. Modulation of Ni⋯Ni separation in synthetic complexes is fascinating in relation to the involvement and stabilization of the dinickel motif in the catalytic transition state of urease. Interaction and binding of cationic complexes (1–2b) with human serum albumin (HSA) and calf thymus-DNA have been examined using spectroscopic techniques. Tryptophan fluorescence quenching of HSA by cationic bimetallic complexes (1, 2a and 2b) featuring a hydrophobic ligand environment shows spontaneous and favorable interactions in the order of 2a > 2b > 1. For the interaction study with CT-DNA, nearly planar complex 1 binds more efficiently than 2a and 2b in octahedral coordination geometry.

 Please wait while we load your content...
Please wait while we load your content...