Assessing the performance of the MM/PBSA and MM/GBSA methods. 6. Capability to predict protein–protein binding free energies and re-rank binding poses generated by protein–protein docking†
Abstract
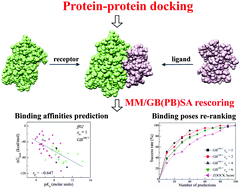
Understanding protein–protein interactions (PPIs) is quite important to elucidate crucial biological processes and even design compounds that interfere with PPIs with pharmaceutical significance. Protein–protein docking can afford the atomic structural details of protein–protein complexes, but the accurate prediction of the three-dimensional structures for protein–protein systems is still notoriously difficult due in part to the lack of an ideal scoring function for protein–protein docking. Compared with most scoring functions used in protein–protein docking, the Molecular Mechanics/Generalized Born Surface Area (MM/GBSA) and Molecular Mechanics/Poisson Boltzmann Surface Area (MM/PBSA) methodologies are more theoretically rigorous, but their overall performance for the predictions of binding affinities and binding poses for protein–protein systems has not been systematically evaluated. In this study, we first evaluated the performance of MM/PBSA and MM/GBSA to predict the binding affinities for 46 protein–protein complexes. On the whole, different force fields, solvation models, and interior dielectric constants have obvious impacts on the prediction accuracy of MM/GBSA and MM/PBSA. The MM/GBSA calculations based on the ff02 force field, the GB model developed by Onufriev et al. and a low interior dielectric constant (εin = 1) yield the best correlation between the predicted binding affinities and the experimental data (rp = −0.647), which is better than MM/PBSA (rp = −0.523) and a number of empirical scoring functions used in protein–protein docking (rp = −0.141 to −0.529). Then, we examined the capability of MM/GBSA to identify the possible near-native binding structures from the decoys generated by ZDOCK for 43 protein–protein systems. The results illustrate that the MM/GBSA rescoring has better capability to distinguish the correct binding structures from the decoys than the ZDOCK scoring. Besides, the optimal interior dielectric constant of MM/GBSA for re-ranking docking poses may be determined by analyzing the characteristics of protein–protein binding interfaces. Considering the relatively high prediction accuracy and low computational cost, MM/GBSA may be a good choice for predicting the binding affinities and identifying correct binding structures for protein–protein systems.


 Please wait while we load your content...
Please wait while we load your content...