Mononuclear Cu(ii) and Zn(ii) complexes with a simple diamine ligand: synthesis, structure, phosphodiester binding and DNA cleavage studies†
Abstract
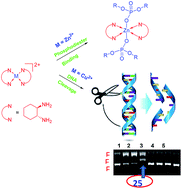
Two mononuclear complexes, [Cu(dach)2(ClO4)2] 1a and [Zn(dach)2](ClO4)2 2a, with simple 1,2-diaminocyclohexane (dach) ligands were synthesized and characterized by elemental analysis, UV-Vis, EPR and IR spectroscopy. The single crystal XRD of 1a and 2a shows that the metal ion has octahedral and tetrahedral geometry, respectively. Both complexes bind with phosphodiesters such as BNPP (bis(p-nitrophenyl)phosphate) and DPP (diphenylphosphate) and their binding affinity was studied by electronic absorption spectroscopy. Complex 1a binds with DPP with Kb = 23 M−1 and 2a shows Kb = 1.15 × 103 M−1. Both complexes interact with BNPP to result in [Cu(dach)2(BNPP)2] 3, [Zn(dach)2(BNPP)2] 4, respectively. These phosphate adducts were well characterized by different spectroscopic techniques and single crystal XRD. The X-ray structures confirm that complexes 3 and 4 are mononuclear with Cu(II) and Zn(II) in octahedral geometry provided by two bidentate dach ligands and two monodentate O donors from BNPP moieties. The nuclease activity of 1a and 2a with supercoiled plasmid DNA pBR322 was examined by gel electrophoresis, which reveals that complex 1a exhibits a significant DNA cleavage activity via oxidative mechanism whereas the corresponding complex 2a did not cleave the DNA. To understand the stereo chemical influence of the dach ligand on DNA cleavage activity, copper and zinc complexes derived from enantiopure form of dach namely [Cu((R,R)-trans-dach)2(ClO4)2] 1b, [Cu((S,S)-trans-dach)2(ClO4)2] 1c & [Cu(cis-dach)2(ClO4)2] 1d and [Zn((R,R)-trans-dach)2](ClO4)2 2b, [Zn((S,S)-trans-dach)2](ClO4)2 2c & [Zn(cis-dach)2](ClO4)2 2d were also investigated. All the Cu(II) complexes oxidatively cleave pBR322 at 25 µM, out of which complex 1b is remarkable to cleave up to 20% Form II and 80% Form III. The binding affinity of these complexes with CT-DNA (Calf thymus-DNA) was studied by electronic absorption spectroscopy and showed moderate to strong Kb values. Electrochemical analyses, viscosity measurements and circular dichroism (CD) studies were carriedout to understand the mode of binding with CT-DNA. The shift in ΔEp, E1/2 and Ipc values suggest that there is a groove interaction existing between CT-DNA and copper complexes. The groove binding is further supported by viscosity measurements and CD spectra.

 Please wait while we load your content...
Please wait while we load your content...