Detection of sequence-specific DNA with a morpholino-functionalized silicon chip
Abstract
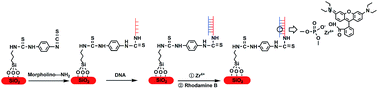
In this work, an efficient method for the sequence-specific detection of DNA based on a morpholino-functionalized silicon chip platform has been proposed. Briefly, morpholino was first immobilized on the surface of a silicon chip using 3-aminopropyltriethoxysilane (APTES) as the silane coupling agent and 1,4-phenylenediisothiocyanate (PDITC) as the cross-linker and then hybridized with DNA in the ensuing step. The fluorescence label was introduced by strongly binding Rhodamine B, which contains a terminal carboxylic group, with DNA by means of phosphate–zirconium–carboxylate coordination reaction. X-ray photoelectron spectroscopy (XPS) was used to characterize the silicon surface. Under optimal conditions, the morpholino-functionalized silicon chip presented a great linear relationship between the fluorescence intensity and the logarithm of target DNA concentrations in the range from 1 pM to 1 nM with a detection limit of 4.52 pM. Furthermore, fully complementary versus single-base mismatched, three-base mismatched and non-complementary DNA could be effectively identified. The chip showed excellent stability because it could be reused for another hybridization experiment after denaturing the morpholino–complementary DNA duplex. In addition, the chip rendered satisfactory analytical performance for the detection of DNA in serum samples, thus exhibiting practical significance. Morpholino-functionalized silicon chips display good sensitivity and selectivity for the detection of DNA and promising applications in single-nucleotide polymorphisms (SNPs).

 Please wait while we load your content...
Please wait while we load your content...