RNA and DNA binding of inert oligonuclear ruthenium(ii) complexes in live eukaryotic cells
Abstract
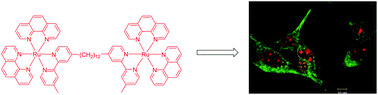
Confocal microscopy was used to study the intracellular localisation of a series of inert polypyridylruthenium(II) complexes with three eukaryotic cells lines – baby hamster kidney (BHK), human embryonic kidney (HEK-293) and liver carcinoma (Hep-G2). Co-staining experiments with the DNA-selective dye DAPI demonstrated that the di-, tri- and tetra-nuclear polypyridylruthenium(II) complexes that are linked by the bis[4(4′-methyl-2,2′-bipyridyl)]-1,12-dodecane bridging ligand (“bb12”) showed a high degree of selectivity for the nucleus of the eukaryotic cells. Additional co-localisation experiments with the general nucleic acid stain SYTO 9 indicated that the ruthenium complexes showed a considerable preference for the RNA-rich nucleolus, rather than chromosomal DNA. No significant differences were observed in the intracellular localisation between the ΔΔ and ΛΛ enantiomers of the dinuclear complex. Cytotoxicity assays carried out over 72 hours indicated that the ruthenium complexes, particularly the tri- and tetra-nuclear species, were significantly toxic to the eukaryotic cells. However, when the activity of the least cytotoxic compound (the ΔΔ enantiomer of the dinuclear species) was determined over a 24 hour period, the results indicated that the ruthenium complex was approximately a 100-fold less toxic to liver and kidney cells than to Gram positive bacteria. Circular dichroism (CD) spectroscopy was used to examine the effect of the ΔΔ and ΛΛ enantiomers of the dinuclear complex on the solution conformations of RNA and DNA. The CD experiments indicated that the RNA maintained the A-type conformation, and the DNA the B-type structure, upon binding by the ruthenium complexes.
- This article is part of the themed collection: Metal Interactions with Nucleic Acids

 Please wait while we load your content...
Please wait while we load your content...