A novel and versatile nanomachine for ultrasensitive and specific detection of microRNAs based on molecular beacon initiated strand displacement amplification coupled with catalytic hairpin assembly with DNAzyme formation†
Abstract
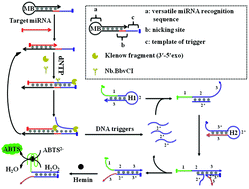
MicroRNAs are small regulatory molecules that can be used as potential biomarkers of clinical diagnosis, and efforts have been directed towards the development of a simple, rapid, and sequence-selective analysis of microRNAs. Here, we report a simple and versatile colorimetric strategy for ultrasensitive and specific determination of microRNAs based on molecular beacon initiated strand displacement amplification (SDA) and catalytic hairpin assembly (CHA) with DNAzyme formation. The presence of target microRNAs triggers strand displacement amplification to release nicking DNA triggers, which initiate CHA to produce large amounts of CHA products. Meanwhile, the numerous CHA products can combine with hemin to form G-quadruplex/hemin DNAzyme, a well-known horseradish peroxidase (HRP) mimic, catalyzing a colorimetric reaction. Moreover, the purification of the SDA mixture has been developed for eliminating matrix interference to decrease nonspecific CHA products. Under the optimal conditions and using the promising amplification strategy, the established colorimetric nanomachine (biosensor) shows high sensitivity and selectivity in a dynamic response range from 5 fM to 5 nM with a detection limit as low as 1.7 fM (S/N = 3). In addition, a versatile colorimetric biosensor has been developed for detection of different miRNAs by only changing the miRNA-recognition domain of molecular beacon. Thus, this colorimetric biosensor may become a potential alternative tool for biomedical research and clinical molecular diagnostics.

 Please wait while we load your content...
Please wait while we load your content...