Internal transcribed spacer rRNA gene sequencing analysis of fungal diversity in Kansas City indoor environments
Abstract
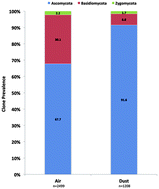
Compared to traditional methods of fungal exposure assessment, molecular methods have provided new insight into the richness of fungal communities present in both indoor and outdoor environments. In this study, we describe the diversity of fungi in the homes of asthmatic children located in Kansas City. Fungal diversity was determined by sequencing the internal transcribed spacer (ITS) regions of ribosomal RNA derived from fungi collected in air and dust samples from 31 homes participating in the Kansas City Safe and Healthy Homes Program (KCSHHP). Sequencing results were then compared to data obtained using viable and non-viable fungal exposure assessment methods. ITS clone libraries were predominantly derived from the phylum Ascomycota in both air (68%) and dust (92%) samples and followed by the Basidiomycota and Zygomycota. The majority of Ascomycota clones belonged to four orders including the Pleosporales, Eurotiales, Capnodiales, and Dothideales. ITS sequencing revealed the presence of a number of rarely documented fungal species placed in the Pleosporales. Several species placed in the Basidiomycota were detected in ITS clone libraries but not by viable or non-viable methods. The prevalence of organizational taxonomic units (OTUs) was significantly higher in air than in dust samples (p < 0.0001); however, no differences between OTUs in air samples collected in the subjects' room and basement were observed. These sequencing results demonstrate a much broader diversity of Ascomycota and Basidiomycota communities in KCSHHP indoor environments than previously estimated using traditional methods of assessment.

 Please wait while we load your content...
Please wait while we load your content...