Hyperswarming adaptations in a bacterium improve collective motility without enhancing single cell motility†
Abstract
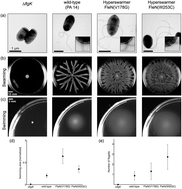
Pseudomonas aeruginosa is a monoflagellated bacterium that can use its single polar flagellum to swim through liquids and move collectively over semisolid surfaces, a behavior called swarming. Previous studies have shown that experimental evolution in swarming colonies leads to the selection of hyperswarming bacteria with multiple flagella. Here we show that the advantage of such hyperswarmer mutants cannot be explained simply by an increase in the raw swimming speed of individual bacteria in liquids. Cell tracking of time-lapse microscopy to quantify single-cell swimming patterns reveals that both wild-type and hyperswarmers alternate between forward and backward runs, rather than doing the run-and-tumble characteristic of enteric bacteria such as E. coli. High-throughput measurement of swimming speeds reveals that hyperswarmers do not swim faster than wild-type in liquid. Wild-type reverses swimming direction in sharp turns without a significant impact on its speed, whereas multiflagellated hyperswarmers tend to alternate fast and slow runs and have wider turning angles. Nonetheless, macroscopic measurement of swimming and swarming speed in colonies shows that hyperswarmers expand faster than wild-type on surfaces and through soft agar matrices. A mathematical model explains how wider turning angles lead to faster spreading when swimming through agar. Our study describes for the first time the swimming patterns in multiflagellated P. aeruginosa mutants and reveals that collective and individual motility in bacteria are not necessarily correlated. Understanding bacterial adaptations to surface motility, such as hyperswarming, requires a collective behavior approach.
- This article is part of the themed collection: Proteins, cells, and tissues in patterned environments

 Please wait while we load your content...
Please wait while we load your content...