Identification and discrimination of binding sites of an organoruthenium anticancer complex to single-stranded oligonucleotides by mass spectrometry†
Abstract
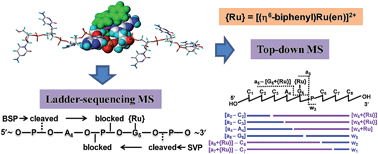
We here report the identification of the binding sites of an organometallic ruthenium anticancer complex [(η6-biphenyl)Ru(en)Cl]+ (1) to single-stranded oligodeoxynucleotides (ODNs) 5′-CCCA4G5C6CC-3′ (I) and 5′-CCC3G4A5CCC-3′ (II) by mass spectrometry. The MS analysis of exonuclease ladders demonstrated that the 5′-exonuclease bovine spleen phosphodiesterase digestion of I and II mono-ruthenated by complex 1 was arrested solely at A4 and partially at C3 and G4, respectively, and that the 3′-exonuclease snake venom phosphodiesterase digestion of the ruthenated ODNs was arrested solely at G5 and G4, respectively, due to the ruthenation. These results did not allow unambiguous identification of ruthenation sites on the metallated ODNs. In contrast, tandem mass spectrometry analysis with CID fragmentation of the mono-ruthenated ODNs provided sequential and complementary [ai − B]/wi fragments, leading to unambiguous identification of G5 in I and G4 in II as the ruthenation sites on the ODN adducts, which is in line with the high selectivity of the complex towards guanine base as reported previously. These findings suggest that caution should be raised with regard to the identification of the binding sites of metal complexes, in particular complexes with bulky ligands, like biphenyl in complex 1, to DNA by MS analysis of exonuclease ladders of the metallated adducts, because the bulky ligands may adopt such an orientation that they block the exonuclease cleavage of the 5′- or 3′-side phosphodiester bonds adjacent to the binding sites, leading to digestion stalling at the nucleotides before the binding sites.

 Please wait while we load your content...
Please wait while we load your content...