Structural and energetic basis for hybridization limits in high-density DNA monolayers†
Abstract
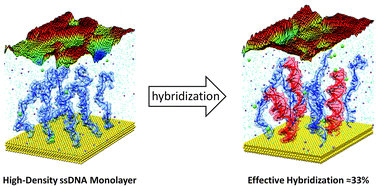
High-density monolayers (HDMs) of single-strand (ss) DNA are important nanoscale platforms for the fabrication of sensors and for mechanistic studies of enzymes on surfaces. Such systems can be used, for example, to monitor gene expression, and for the construction of more complex nanodevices via selective hybridization with the complementary oligos dissolved in solution. In this framework, controlling HDM hybridization is essential to control the final properties. Different studies demonstrate that at the typical density of ≈1013 molecules per cm2 no more than ≈30–40% of the HDM ssDNA is successfully hybridized. Until now, however, the origin of the HDM hybridization limit has remained unclear. In this work, molecular dynamics (MD) simulations of HDM systems with variable hybridization reveal that, independently of other experimental parameters, the effective hybridization for a HDM of this density is intrinsically limited by molecular and electrostatic crowding. A detailed structural analysis of the HDM model shows good agreement with our

 Please wait while we load your content...
Please wait while we load your content...