Macromolecular platinum-drugs based on statistical and block copolymer structures and their DNA binding ability†
Abstract
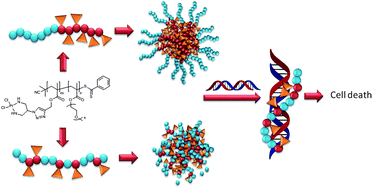
Macromolecular platinum drugs were prepared from statistical copolymers or block copolymers. Reversible addition–fragmentation chain-transfer (RAFT) polymerisation was applied to obtain copolymers based on 3-(trimethylsilyl) prop-2-ynyl methacrylate (TMSPMA) as the reactive building block for the conjugation of platinum drugs and oligo(ethylene glycol methyl ether) methacrylate (OEGMEMA) as the water-soluble part. Two statistical copolymers and one block copolymer were obtained, which were deprotected to allow the subsequent ‘click’ reaction. Inspired by the platinum anti-cancer drug cisplatin (cis-diamminedichloroplatinum(II)), the platinum drug was conjugated to the polymer via a stable, permanent amino ligand. Therefore, a BOC protected 2-azidopropane-1,3-diamine (N3-DAP-BOC) ligand was prepared, which was attached to the polymers by copper catalysed azide–alkyne Huisgen cycloaddition ‘click’ (CuACC) reaction. Purification of the polymer to remove the copper complex was found to be difficult and several approaches were investigated. Subsequent platinum drug conjugation led to three different macromolecular platinum drugs P(OEGMEMA64-co-[(PMA-DAP)8-Pt7]), P(OEGMEMA62-co-[(PMA-DAP)16-Pt14]), and P(OEGMEMA84-b-[(PMA-DAP)48-Pt20]) with conjugation efficiencies of more than 80% for the statistical copolymers and 40% for the block copolymer. Binding efficiency to DNA of all three polymers was studied using fluorescence spectroscopy revealing the highest binding with the statistical copolymer with the highest drug density. Interestingly, the toxicity of all three polymers towards the ovarian cancer cell line OVCAR-3 highlights the importance of further work that investigates cellular uptake and stability of the aggregates in vitro.

 Please wait while we load your content...
Please wait while we load your content...