Molecular recognition between DNA and a copper-based anticancer complex†
Abstract
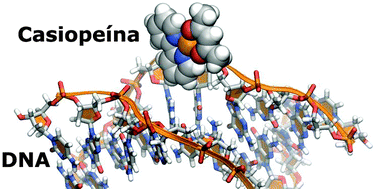
The aim of this work is to describe the specific recognition site between DNA and an anticancer copper complex by means of computational methods. Molecular dynamics were used to find the preferred site of binding between selected DNA chains and [Cu(2,2′-bipyridine)(acetylacetonate)(H2O)]+ (Cas). Full DFT optimizations of selected geometries extracted from simulations, followed by a topological analysis of electron density allowed us to define the specific interactions inside the recognition site. Cas links deoxyribose-phosphate by a coordination bond between metal and the

 Please wait while we load your content...
Please wait while we load your content...